As a scientist I have had the incredible opportunity to travel the world, contributing to research projects in beautiful places with fantastic mentors.
HOPKINS MARINE STATION
My PhD was completed in Lowe Lab at the Hopkin’s Marine Station.
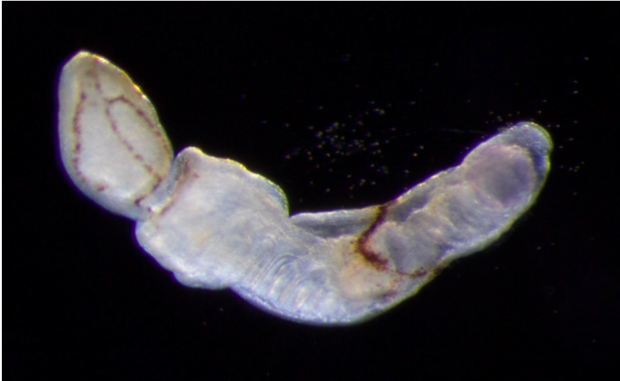
My thesis work involved studying an indirect developing enteropneust hemichordate worm Schizocardium californicum with distinct larval and adult body plans. While which the most common developmental strategy in many animal phyla, much of what we know about development comes from direct-developing species. Across metazoan diversity, some indirect developing animals remodel and transform an existing larval body into the adult body plan. Many questions remain in regard to the general mechanisms of these processes. Do the larval cells of indirect developing animals have a distinct larval fate or trajectory? In this scheme of indirect development, larval-specific cells must be eliminated, and new cells must proliferate to form the adult body plan. Or, is it possible that larval cell types retain developmental plasticity and have the capacity to differentiate into the appropriate cells of the adult body plan? In an animal that transforms an existing body from larva to adult, these transitions provide a window into tissue turnover and remodeling, an important focus of cell and developmental biology.
Publications:
Bump P, Khariton M, Stubbert C, Moyen NE, Yan J, Wang B, Lowe CJ. Comparisons of cell proliferation and cell death from tornaria larva to juvenile worm in the hemichordate Schizocardium californicum. EvoDevo. 2022 Dec;13(1):13.
Bump P, Lubeck L. Marine Invertebrates One Cell at A Time: Insights from Single-Cell Analysis. Integrative And Comparative Biology. 2023 May 15;icad034. PMID: 37188638
Lin CY, Marlétaz F, Pérez-Posada A, Martínez-García PM, Schloissnig S, Peluso P, Conception GT, Bump P, Chen YC, Chou C, Lin CY, Fan TP, Tsai CT, Skarmeta JLG, Tena JJ, Lowe CJ, Rank DR, Rokhsar DS, Yu JK, Su YH. (2024) Chromosome-level genome assemblies of 2 hemichordates provide new insights into deuterostome origin and chromosome evolution. PLOS Biology 22(6): e3002661
BROAD CENTER FOR REGENERATIVE MEDICINE AND STEM CELL RESEARCH
Research Lab Technician
Mentor: Dr. Gage Crump
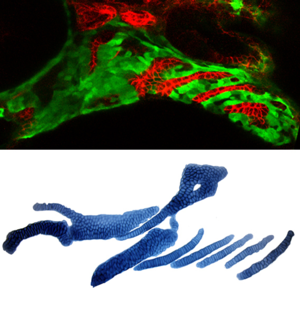
The Crump Lab is interested in how genetic information is decoded to generate the precise three-dimensional shapes of organs. As a model, the lab studies the facial cartilages and bones of the larval zebrafish, which can be continuously imaged over time in living transgenic embryos. Specific research topics include the developmental origins of the neural crest progenitor cells that build the craniofacial skeleton. As an example, the pharyngeal arches, where the neural crest cells arise (to right in green), transform into the adult facial skeleton (to the right in blue).
My work in the lab focused on screening a large number of novel candidate genes whose expression is highly enriched in a population of cells known as the early neural crest.
Publications:
Paudel S, Gjorcheska S, Bump P, Barske L. Patterning of cartilaginous condensations in the developing facial skeleton. Developmental Biology. 2022 Jun 1;486:44-55.
Askary A, Xu P, Barske L, Bay M, Bump P, Balczerski B, Bonaguidi MA, Crump JG. Genome-wide analysis of facial skeletal regionalization in zebrafish. Development. 2017;144(16):2994–3005. PMID: 28705894 PMCID: PMC5592815
Barske L, Askary A, Zuniga E, Balczerski B, Bump P, Nichols JT, Crump JG. Competition between Jagged-Notch and Endothelin1 Signaling Selectively Restricts Cartilage Formation in the Zebrafish Upper Face. Mullins MC, editor. PLoS Genet. 2016 Apr 8;12(4):e1005967
MARINE BIOLOGICAL LABORATORY

Biological Discovery in Woods Hole Summer Opportunity for Undergraduate Research
The Synergistic Nature of the Behaviors and Mechanisms that Support Effective Burrowing in the Mantis Shrimp Squilla empusa.
Mentor: Dr. Kristina Mead-Vetter
The mantis shrimp Squilla empusa is a charismatic marine crustacean known for its powerful strike, keen sense of vision, and chemosensory abilities. These benthic creatures create extensive burrows that are important in feeding, reproduction, and protection from predation. Through field observations of a population located in Great Harbor in Woods Hole, MA this species of mantis shrimp has been observed to construct burrows faster and makes more alterations than previously recorded. To understand the mechanics of these burrowing behaviors, mantis shrimp were filmed making burrows in the lab using high-speed videography. S. empusa used two markedly distinct methods of burrowing: pleopod fanning and maxilliped bulldozing. Through this series of observations and analyses we are starting to understand how pleopod anatomy and kinematics work synergistically to create an effective burrowing system.
BERMUDA INSTITUTE OF OCEAN SCIENCES

Bermuda Institute of Ocean Sciences REU.
Mentor: Dr. Andrea Bodnar
Project: Molecular Profile of Stem Cells in Sea Urchin Tissues.
This project investigated the temporal and spatial expression of a panel of stem cell genes in embryos and adult tissues of sea urchins. These studies will involve both quantitative reverse transcriptase PCR and RNA in situ hybridization.The ultimate goal of this research is to begin to understand the role that stem cells play in tissue maintenance and regeneration and how this impacts the life histories of these animals.
KEWALO MARINE LABORATORY

EDEN Undergraduate Internship, Undergraduate Honors Thesis
Mentor: Dr. Michael Hadfield
Project: Transcriptomes of Pre-competent, Competent, and Metamorphosed Larvae of Hydrodies elegans Reveal the Distinct Role of Innate Immunity Genes in Response to a Bacterial Settlement Cue.
Hydroides elegans is a common serpulid polychaete, which like all invertebrates, lacks an antibody-based adaptive immunity. Invertebrates have evolved efficient defense responses to pathogens as well as sophisticated systems for recognizing and differentiating beneficial and pathogenic microorganisms. An important factor associated with invertebrate innate immunity, are microorganism-associated molecular patterns (MAMPs), which can initiate signaling cascades by first interacting with pattern recognition receptors (PRRs) such as Toll-like receptors (TLRs) and peptidoglycan recognition proteins (PGRPs). Both MAMP-mediated signaling can initiate signaling cascades, usually through the nuclear factor-κB (NF-κB) or similar pathway, that lead to the activation of a variety of immune responses. The concept of an immune response to MAMPs is particularly relevant for H. elegans larvae, which are induced to settle and rapidly metamorphose by the presence of a well-developed bacterial biofilm. The ultimate goal of this work is to elucidate the importance of differentially expressed innate immunity genes. This will enhance our understanding of the role of these genes in responding to cues from marine bacteria, as well as identifying gene function and pathways in the metamorphosis.